The Sentinel Surveillance Network identified 11 cases of influenza-like illness, thus exceeding the recommended threshold for the new epidemic season, according to the European Center for Disease Prevention and Control (ECDC) guidelines.
Regarding SARS-CoV-2 genomic surveillance, the LNS analysed 748 specimens from residents in Luxembourg in week 47/2021 (from 2373 total cases in the Grand Duchy of Luxembourg, 31,5%). This does not reach the new ECDC recommendations to detect emerging variants at 1% prevalence (minimum sample size of 927). Complementing the above sequencing, 387 additional specimens from residents were screened for the Omicron variant, to increase our detection capacity.
All specimens were assigned to the Delta variant, no Omicron cases were detected. Community surveillance showed that the AY.43 lineage continues to be the most frequent one (35,1%), followed by AY.4 (10,4%) and AY.98.1 (8,1%). Regarding target groups, the same three lineages were found to be the most frequent ones, both within hospital specimens and post-vaccination breakthrough cases. As for the mutations under surveillance, they revealed no outstanding behaviour, remaining in agreement with the lineages observed.

The Laboratoire national de santé, as National Reference Laboratory for Acute Respiratory Infections in Luxembourg, performs close surveillance on respiratory viruses, with a special focus on SARS-CoV-2. There are currently two active projects on which the ReViLux provides updates:
The Sentinel Surveillance Network. It provides a broad picture of respiratory diseases affecting the Luxembourgish population, based on its double monitoring system (syndromic and viroogical).
The National SARS-COV-2 Genomic Surveillance Program. It enables detailed observation of SARS-CoV-2 mutations and variants through time and space, and also monitoring specific groups of interest.
The Sentinel Surveillance Network aims at monitoring the circulating respiratory viruses, including SARS-CoV-2, and hence underpin public health actions. Following the World Health Organization (WHO) and European Centre for Disease Prevention and Control (ECDC) guidance, it focuses on cases of acute respiratory infection (ARI) and influenza-like illness (ILI).
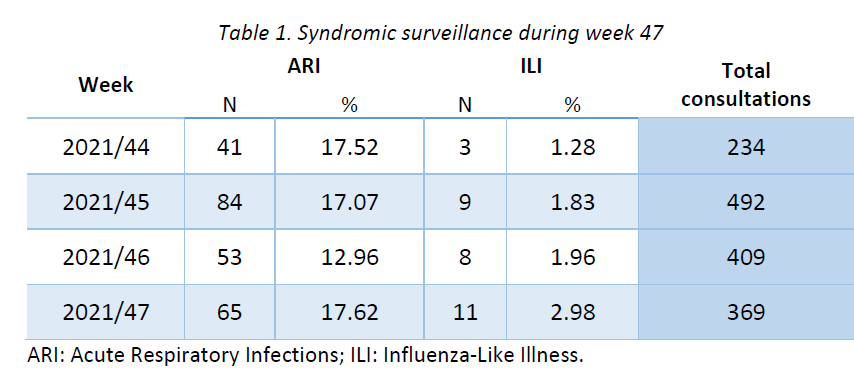
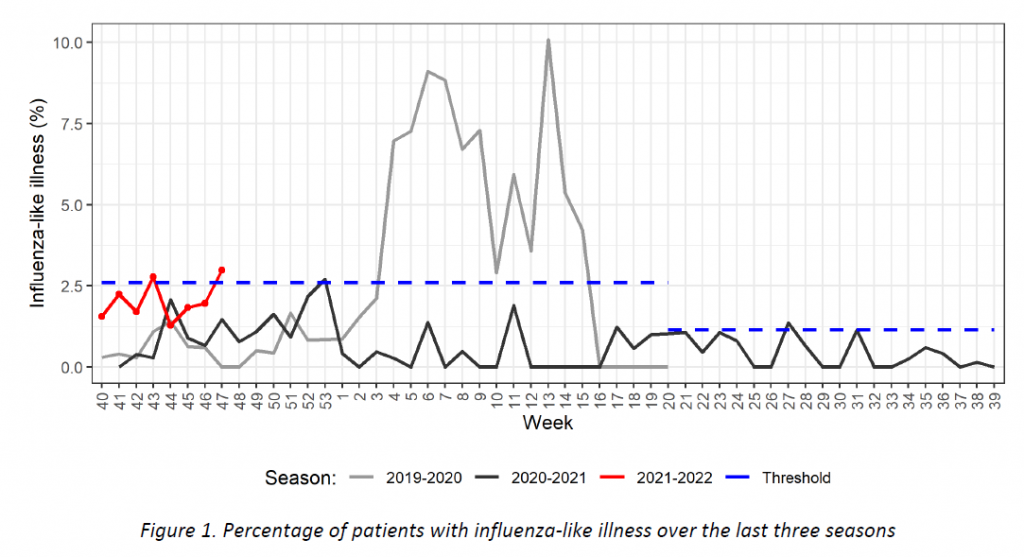
Week 40 marked the beginning of the new influenza season 2021-2022. Results of syndromic surveillance during the last four weeks are displayed in Table 1 and the history of ILI consultations since the 2019-2020 season is shown in Figure 1. Eleven cases of ILI were identified in week 47 (out of 369 consultations); therefore, the percentage of ILI (2,98%) exceeds the threshold for the epidemic season (2,59%), according to the ECDC.
Regarding the virological surveillance, a partnership among the CNS, private laboratories and the LNS recently started and will enable us to monitor the presence of several respiratory viruses. Results from the first analyses will be published soon.
The National Reference Laboratory for Acute Respiratory Infections at LNS receives SARS-CoV-2 positive samples (nasopharyngeal or oropharyngeal swabs analysed by RT-PCR) from the national network of laboratories and proceeds as follows:
Sequencing all specimens from hospital cases.
Sequencing all specimens from post-vaccination cases.
Sequencing specimens from clusters with high transmission.
Sequencing a representative sample of community cases.
The representative sample of community cases is a systematic selection from all SARS-CoV-2 positive cases registered in Luxembourg to detect emerging variants and early increases in their incidence and transmission within the community in Luxembourg. This sample is selected according to the ECDC guidelines.
Due to the emergence of the new Omicron variant of concern, as well as the high incidence rates in the European context, the LNS is currently carrying out a complementary PCR screening strategy, to enable faster detection and identification of any Omicron variant cases.
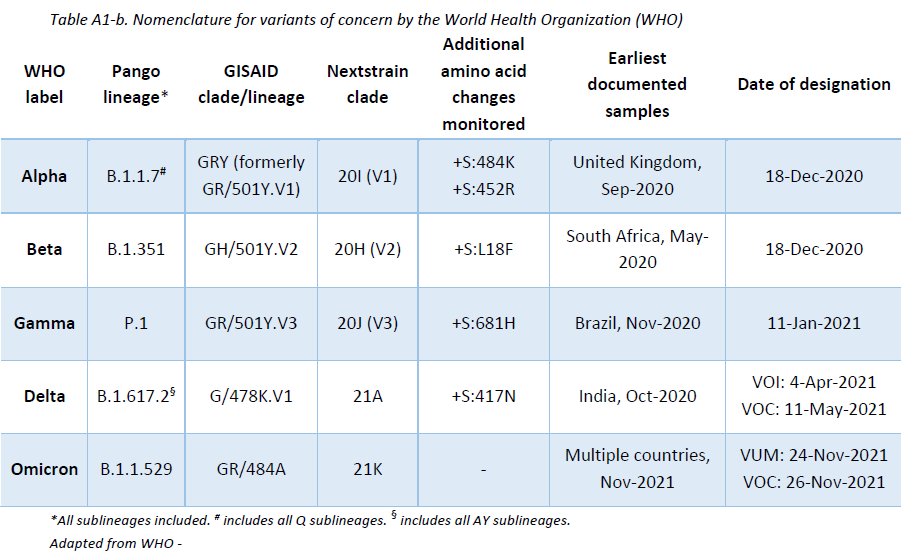
The LNS shares its sequencing results with GISAID EpiCov database periodically. SARS-CoV-2 lineages have been assigned based on Rambaut et al. using the Phylogenetic Assignment of Named Global Outbreak LINeages (pangolin) software (v3.1.16, pangoLEARN 2021-11-25). The Pango nomenclature is used in addition to the WHO nomenclature to enable easier visualization of links between any evolving variants and their ancestor (See nomenclature equivalences in Appendix 1). Delta lineages nomenclature is in constant review. The original Delta B.1.617.2 lineage is being re-classified into more specific AY lineages in order to enable a more precise tracing of the cases. This report is based on the latest nomenclature, and previously assigned lineages might have been updated to remain consistent with the latest nomenclature.
In week 47, 2373 new cases were registered in Luxembourg. Given the epidemiological situation after the emergence of Omicron, achieving a 1% incidence detection power is preferrable than the usual 2.5%, according to the ECDC. For this reason, the recommended sample size is estimated to be 927 specimens (39,1%).
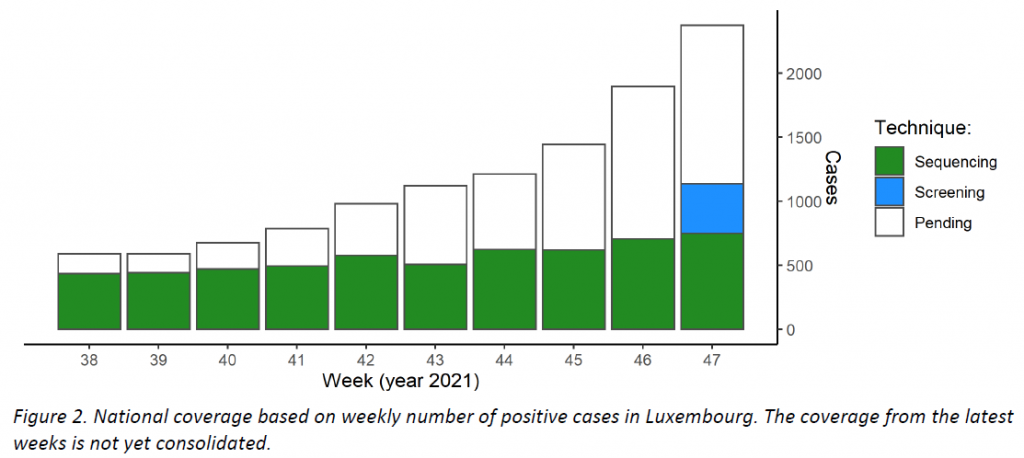
Last week, the microbial genomics unit at the LNS analysed 889 specimens from week 47, with 747 specimens having been collected in week 47 from residents. As this sample size did not reach the ECDC-recommended sample size, another 387 specimens from residents were tested by targeted PCR for the Omicron variant (overall, 47,8% coverage of the 2373 total cases registered in Luxembourg; see coverage trend in Figure 2).
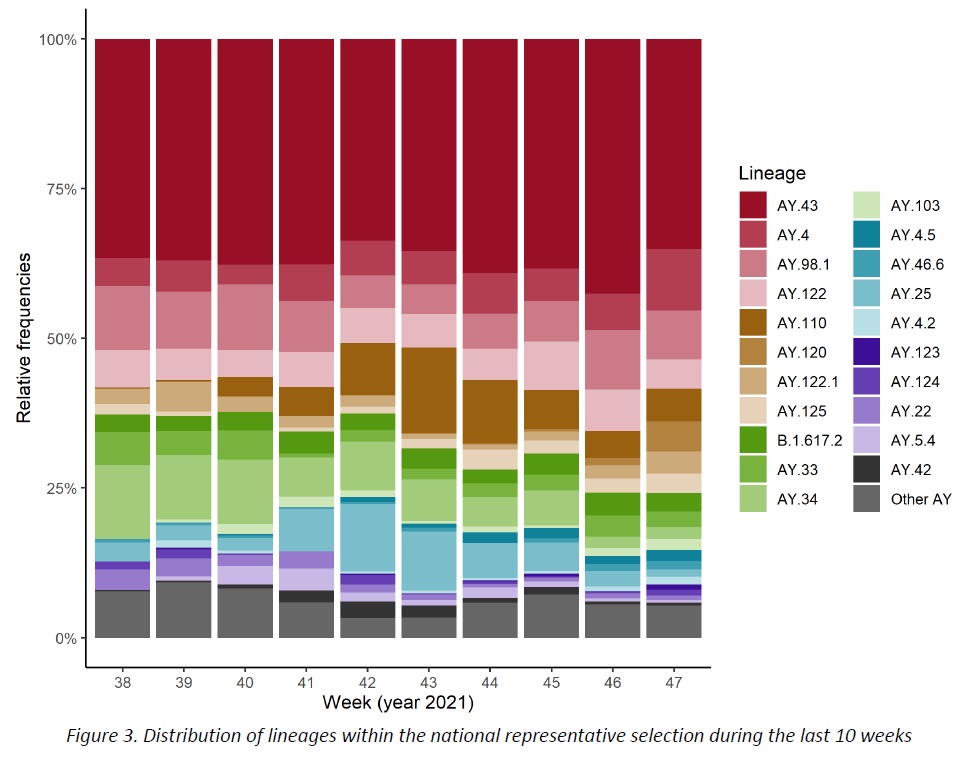
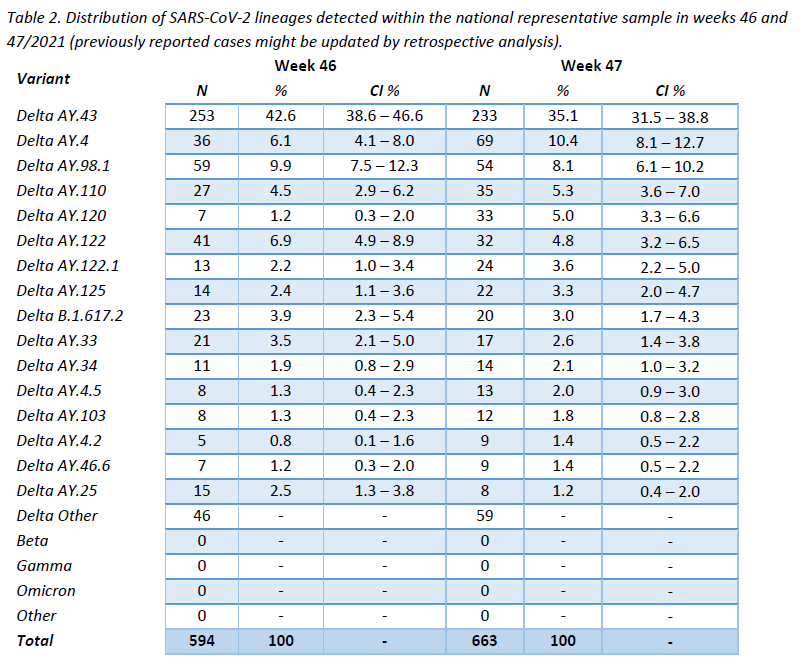
The distribution of successfully assigned lineages within the representative sample is shown in Figure 3. The selection of lineages displayed is based on their pevalence during the last 10 weeks (min. 1%). This distribution is further detailed for the last 2 weeks in Table 2. The lineage AY.43 continues to be the most frequent one (35,1%), followed by AY.4 (10,4%), while the remaining lineages show frequencies lower than 10%.
As for the PCR screening strategy, no specimens were suspicious for the Omicron variant.
In addition to the surveillance of SARS-CoV-2 variants, the LNS monitors the occurrence of SARS-CoV-2 mutations reported to have a clinical and epidemiological relevance. This complementary surveillance enables us to detect unexpected mutations among the specimens sequenced. It is expected that VOC defining mutations share the same distribution as their corresponding VOCs. However, newly acquired mutations may occur and their early detection might be key to expect changes in the epidemic evolution.
Following ECDC guidance, the LNS is currently monitoring 42 mutations to the spike protein frequently associated to VOCs. Accordingly to the distribution lineages, most of the mutations found during the last 2 weeks are linked to Delta cases. No specimen displayed a mutation pattern suspicious of the Omicron variant.
Centers for Disease Control and Prevention. SARS-CoV-2 Variant Classifications and Definitions. Retrieved 6 December 2021, from https://www.cdc.gov/coronavirus/2019-ncov/variants/variant-info.html
COVID-19 Data Portal – accelerating scientific research through data. (2021). Retrieved 6 December 2021, from https://www.covid19dataportal.org/sequences
European Centre for Disease Prevention and Control. Guidance for representative and targeted genomic SARS-CoV-2 monitoring – 3 May 2021. ECDC : Stockholm ; 2021
European Centre for Disease Prevention and Control. SARS-CoV-2 variants of concern. Retrieved 6 December 2021, from https://www.ecdc.europa.eu/en/covid-19/variants-concern
European Centre for Disease Prevention and Control. Epidemiological update: Omicron variant of concern (VOC) – data as of 6 December 2021 (12.00). Retrieved 6 December 2021, from https://www.ecdc.europa.eu/en/news-events/epidemiological-update-omicron-variant-concern-voc-data-6-december-2021
Genomic sequencing of SARS-CoV-2: a guide to implementation for maximum impact on public health. Geneva: World Health Organization; 2021.
GitHub – cov-lineages/pangolin: Software package for assigning SARS-CoV-2 genome sequences to global lineages. (2021). Retrieved 6 December 2021, from https://github.com/cov-lineages/pangolin
Hadfield J., Megill C., Bell S., Huddleston J., Potter B., Callender C. et al. (2018). Nextstrain: real-time tracking of pathogen evolution. Bioinformatics, 34(23), 4121-4123. doi: 10.1093/bioinformatics/bty407
Instituto Nacional de Saúde Doutor Ricardo Jorge. Diversidade genética do novo coronavírus SARS-CoV-2 (COVID-19) em Portugal. Retrieved 6 December 2021, from https://insaflu.insa.pt/covid19/
Pango Network. New AY lineages. Retrieved 6 December 2021, from: https://www.pango.network/new-ay-lineages/
Pango Network. New AY lineages. Retrieved 6 December 2021, from: https://www.pango.network/new-ay-lineages-and-an-update-to-ay-4-ay-12/
Rambaut A., Holmes E., O’Toole Á., Hill V., McCrone J., Ruis C. et al. (2020). A dynamic nomenclature proposal for SARS-CoV-2 lineages to assist genomic epidemiology. Nature Microbiology, 5(11), 1403-1407. doi: 10.1038/s41564-020-0770-5
According to the ECDC

References:
According to the WHO