
The aim of the “Sentinel” national surveillance program is to monitor the circulating respiratory viruses, including SARS-CoV-2 variants, and hence underpin public health actions.
In week 18/2021, the overall frequency of the SARS-CoV-2 B.1.1.7 variant in all sequenced specimens remained stable on 81,1% (CI 77,6% – 84,6%, p<0,05). For the SARS-CoV-2 B.1.351 variant, we found an overall frequency of 6,8% (CI 4,5% – 9%, p<0,05) within the sequenced sample, while for P.1, the frequency increased to 4,6% (CI 2,7% – 6,5%, p<0,05).
The representative sample was estimated, based on the number of positive cases in Luxembourg for week 18 (936). The minimum sample size required to detect prevalence of B.1.1.7 (81%) reported in week 17, with an error margin of 5%, was estimated to be 189 specimens. This number corresponds to a coverage of 20,2 %, which exceeds the minimum coverage recommended by ECDC (10%). The sequencing results of week 16 are representative of the circulating variants in Luxembourg with a margin of error of 5%.
The total number of sequences performed this week was 500, with 482 specimens (deduplicated) having been collected in the time frame of week 18/2021. The sequencing coverage this week was 47,2% from all positive cases in Luxembourg.
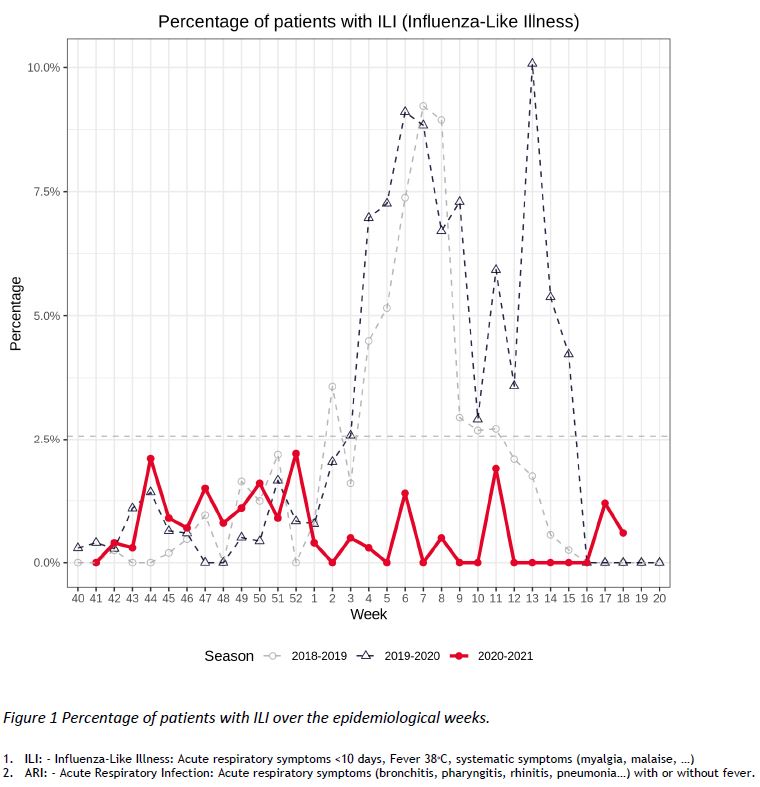
The “Sentinel” surveillance network reported 346 consultations in week 18 (03/MAY/2021 – 09/MAY/2021). There were 2 cases of ILI1, corresponding to 0,6% of the consultations, as shown in Figure 1. The number of consultations for ARI2 was 47, which represents 13,6% of the consultations.
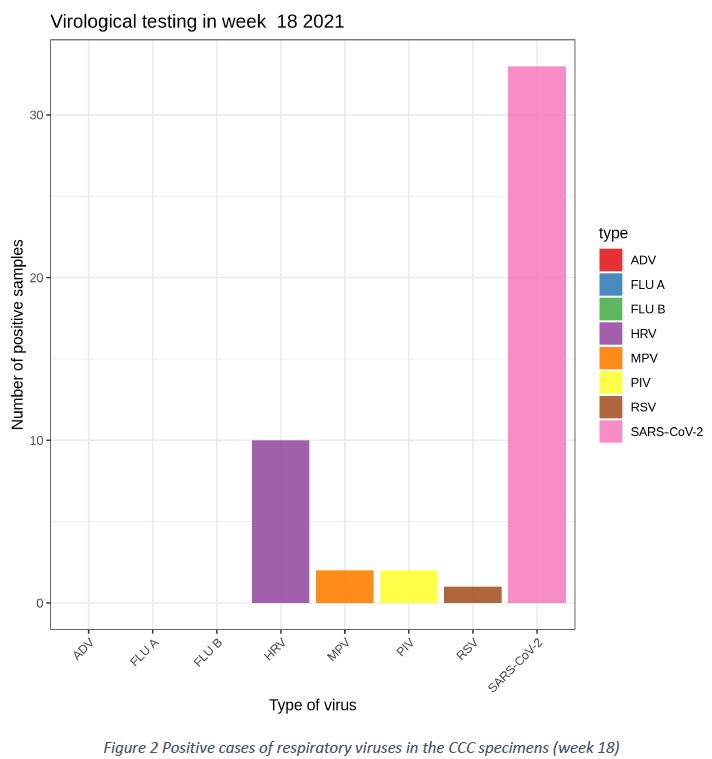
Patients presenting with ILI and ARI at the Covid Consultation Centre (CCC) in Luxembourg were tested using a respiratory virus panel (ADV = Adenovirus, FLU A = Influenza A, FLU B = Influenza B, HRV = Human Rhinovirus, MPV = Human metapneumovirus, PIV = Parainfluenza virus, RSV = Respiratory Syncytial Virus, SARS-CoV-2 = Severe acute respiratory syndrome coronavirus 2).
The SARS-COV-2 was the most prevalent respiratory virus detected in the “Sentinel” network, with 33 positive cases (28,7%). The wave of Human Rhinovirus (HRV) continued in week 18/2021, with 10 positive cases in 115 tests (8,7%). Sporadic cases of other respiratory viruses continued to appear, such as 2 cases of PIV (1,7%) and MPV (each 1,7%) and 1 case of RSV (0,9%). No case of Influenza A/B was detected. (c.f. Figure 2).
In Luxembourg, we have tested 115 samples from the Sentinel surveillance network, as compared to 555 specimens tested in Europe, in the week 18/2021. None of these 555 specimens tested positive for Influenza type A/B virus. Overall, surveillance was very efficient over the course of the 2020-2021 season and although there was a small decrease in the number of samples tested as compared with previous seasons, there was a remarkable decrease (>99% decrease) in the number of influenza infections detected, with numbers detected on a weekly basis being similar to those reported during interseasonal periods. (Source: FluNews Europe).
The National Reference Laboratory for Acute Respiratory Infections at LNS continues to improve the representativeness of the pool of sequenced specimens to reach real-time epidemiology, by implementing the following weekly sequencing activities:
The representative sequencing sample was based on the minimum number of specimens required to extrapolate prevalence of VOC variants with error rate of 5%. The representative sample was estimated based on the number of positive cases in Luxembourg in week 18 (936). The minimum sample size required to detect prevalence of B.1.1.7 (81%) reported in week 17 with an error margin of 5% was estimated to be 189 specimens. The calculation was based on a sample size calculation tool that uses the expected prevalence of the variant in the total population. (Population Proportion – Sample Size – Select Statistical Consultants (select-statistics.co.uk). This number represented a coverage of 20,2% which exceeds the minimum coverage recommended by ECDC (10%). The number of non-targeted specimens from Luxembourgish residents sequenced this week was 397. Therefore, our sequencing results this week are representative of the circulating variants in Luxembourg.
The starting material used for sequencing is respiratory specimens (nasopharyngeal or oropharyngeal swabs) that have already been tested positive by RT PCR.
In parallel, we continue to evaluate commercial PCR screening, using the same starting material, to detect the Variants of Concern (VOCs), with specific PCR. This will enable a faster investigation time of any outbreak and allow us to screen 100% of positive cases referred to LNS, including those that do not pass the quality criteria for sequencing. Specimens that are not eligible for sequencing can thus be used in a PCR to detect the presence of variants of concern.
The LNS sequencing data sharing strategy includes sharing of the sequencing data with GISAID EpiCov database (www.gisaid.org ) on a periodic basis.
Last week the microbial genomics platform at the LNS sequenced 500 specimens, with 482 collected in week 18/2021. The sequencing pool represents 47,2% of new infections reported in Luxembourg in week 18/2021. Among these 482 specimens, 50 specimens were reported to be part of a cluster or outbreak investigation, and 40 specimens were from non-residents (5 specimen overlapping). This leads to 397 specimens, collected in week 18, and being the representative population sequencing sample. In the population representative sample of residents, the frequency of B.1.1.7 and B.1.351 was 83,4% and 5,8% respectively, while the frequency of P.1 was 3,8%.
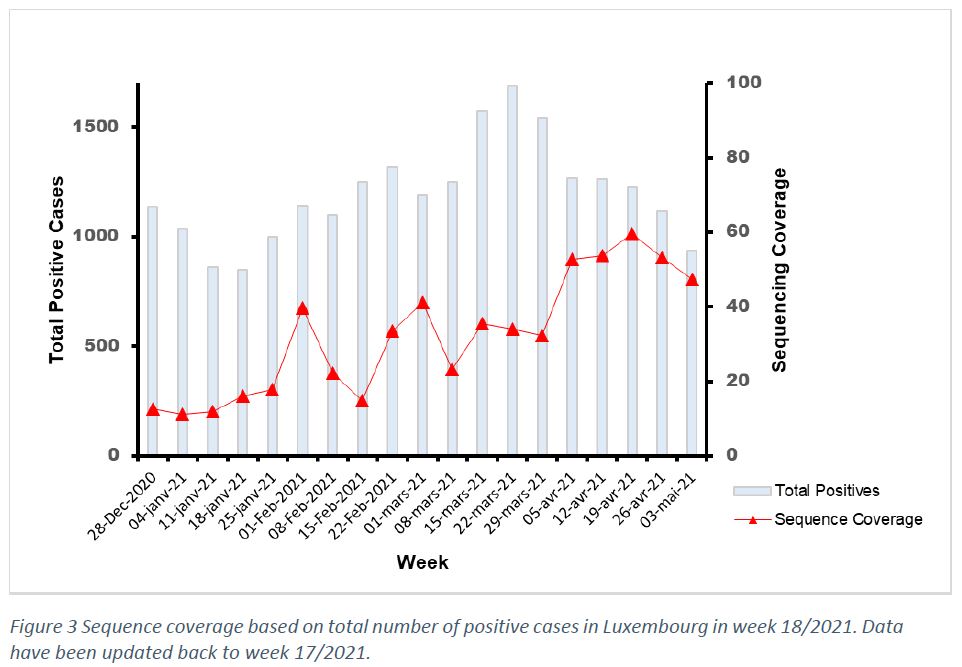
The population sequencing coverage in week 18/2021 was 47,2% (Figure 3). Based on statistical inference, the frequency of the reported variants in week 18/2021 is representative of the circulating variants in Luxembourg with a margin of error of 5%.
Lineages (variants) have been assigned based on Rambaut et al by means of Phylogenetic Assignment of Named Global Outbeak LINeages (pangolin) software (v2.4.2, pangoLEARN version 2021-05-12).
The lineage nomenclature system that we use is the one proposed by Rambaut et al. that focuses on actively circulating virus lineages (https://cov-lineages.org ).
In week 18/2021, in the population representative sample, after removal of cluster specimens, and excluding specimens collected from non-residents, there were 13 circulating SARS-CoV-2 variants, with the main three variants being B.1.1.7 (83,4%, CI 79,7% – 87,1%), B.1.351 (5,8%, CI 3,5% – 8,1%), followed by P.1 (3,8%, CI 1,9% – 5,7%), as shown in Figure 4.
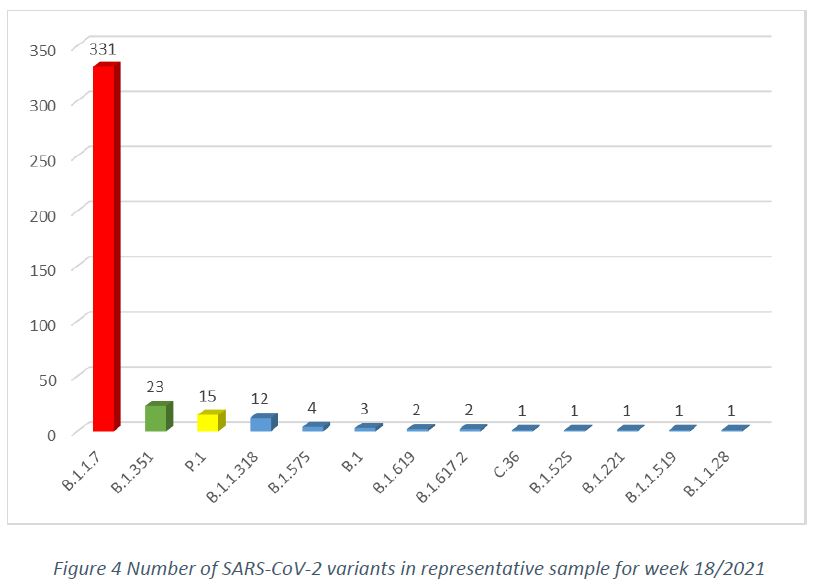
Among specimens collected within the week 18/2021, 391 cases of the B.1.1.7 variant have been detected, representing 81,1% of the specimens in the week’s sequencing pool (by comparison, the week 17/2021 pool had shown a frequency of 80,6% of this variant, including additional specimens having been sequenced from previous weeks). The total case count of sequenced variant B.1.1.7 was 4098 by week 18/2021. The earliest collection date for this variant remains 19/DEC/2020 and the latest is 09/05/2021.
In the collection period of week 18/2021, 33 cases of the South African variant B.1.351 have been detected, representing 6,8% of the specimens in the week’s sequencing pool (by comparison, the week 17/2021 pool had shown 10,8% of this variant, including additional specimens having been sequenced from previous weeks). The total case count of sequenced variant B.1.351 was 967 by week 18/2021. The earliest collection date for this variant remains 11/JAN/2021 and the latest is 09/MAY/2021.
In week 18/2021, 1 additional case of B.1.525 has been detected, and 22 new cases of the Brazilian variant P.1. Thus, the case count by week 18 for B.1.525 is 26 (latest sampling date 07/MAY/2021) and for P.1 the case count is 72 (latest sampling date 08/MAY/2021).
In week 18/2021, 4 additional case of the B.1.617 subtype B.1.617.2 have been detected (latest sampling date 07/MA/2021). The case count by week 18 for B.1.617.1 remains 6, and for B.1.617.2 it raises to 5. Since May 6th B.1.617.2 has been escalated by Public Health Engand from a variant under investigation to a variant of concern.
By now, no case of A.23.1 has been detected in Luxembourg (Figure 5).
Lineage B.1.1.7 is characterized by several spike protein mutations, including N501Y, H69/V70del and P861H. The variant seems to have a considerable epidemiological impact, as it has a higher transmissibility rate.
Lineage B.1.351 holds numerous spike protein mutations, of which three are located in the receptor binding domain (K417N, E484K and N501Y), and are therefore relevant for antibody binding. As for B.1.1.7, a higher transmissibility rate and viral loads seem to be associated with this variant. Due to the K417N and E484K mutations, an impact on vaccination efficacy and possibility of reinfection is subject to scientific investigation.
Lineage P.1 (descendent of B.1.1.28), initially found in the Amazon region, has a similar mutation profile as the South African variant, including E484K and N501Y. Concerns are, as for the South African variant, higher transmissibility and a decreased protection by neutralizing antibodies.
Lineage B.1.525 carries several mutations of biological significance, including E484K, Q677H and F888L. It does not carry N501Y, but a set of deletions similar to the B.1.1.7 variant.
Lineage B.1.617 is a variant first detected in India and was designated “Under Investigation” on 1st April 2021 by Public Health England. It contains a number of spike mutations associated with antigenic escape or found in other variants of concern, including L452R, E484Q and P681R. Subtype B.1.617.2 does not carry S:E484Q and seems to be at least as transmissible as B.1.1.7 (with moderate confidence).
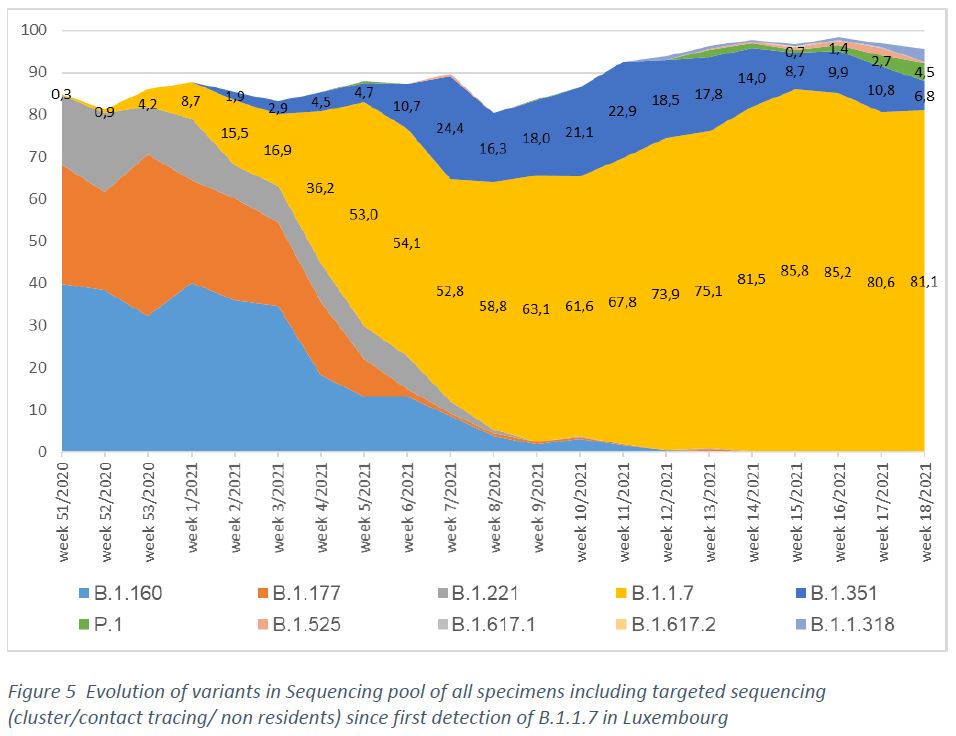
By week 18/2021, 96,5 % of the variants detected in the sequenced specimen pool are declared as either Variants of Concern (VOC – B.1.1.7, B.1.351, P.1, B.1.617.2) or Variants Under Investigation (VUI – B.1.525, B.1.1.318, B.1.617.1).
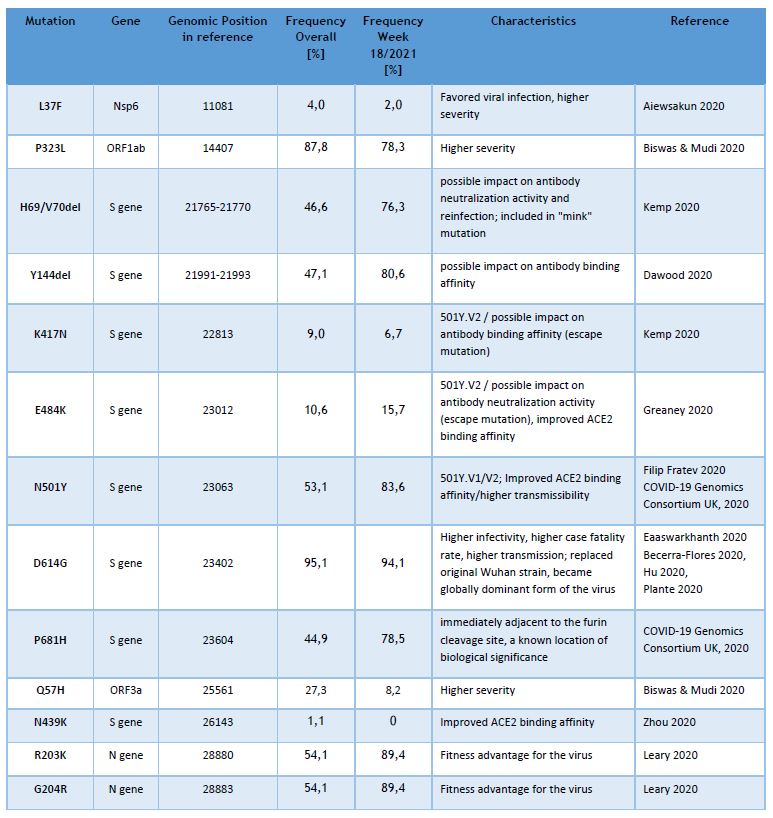
Currently the LNS genomic surveillance program – independently from lineage calling – notes the occurrence of 13 different known SARS-CoV-2 mutations, assumed to have clinical and epidemiological relevance. The list of observed mutations is being updated continually, based on the appearance and prevalence of SARS-CoV-2 variants.
The following table provides the overall frequencies of these mutations, detected in the lineage-assignable genome sequences, analyzed between 01/SEP/2020 and 09/MAY/2021 (N=10831), as well as the frequencies in week 18/2021.
Genomic sequencing of SARS-CoV-2. A guide to implementation for maximum impact on public health. WHO, 8 January 2021.
COVID-19 data portal. 2020 (https://www.covid19dataportal.org/sequences )
J Hadfield et al. Nextstrain: real-time tracking of pathogen evolution. Bioinformatics 2018;34:4121-4123
A Rambaut et al. A dynamic nomenclature proposal for SARS-CoV-2 lineages to assist genomic epidemiology. Nat Microbiol 2020;5:1403-1407
https://github.com/cov-lineages/pangolin
For more information on lineages visit: https://cov-lineages.org
For more information and statistics on Covid-19 infections in Luxembourg visit: https://covid19.public.lu/en.html